Multi-scale spiking network model of macaque visual cortex: Simulation data
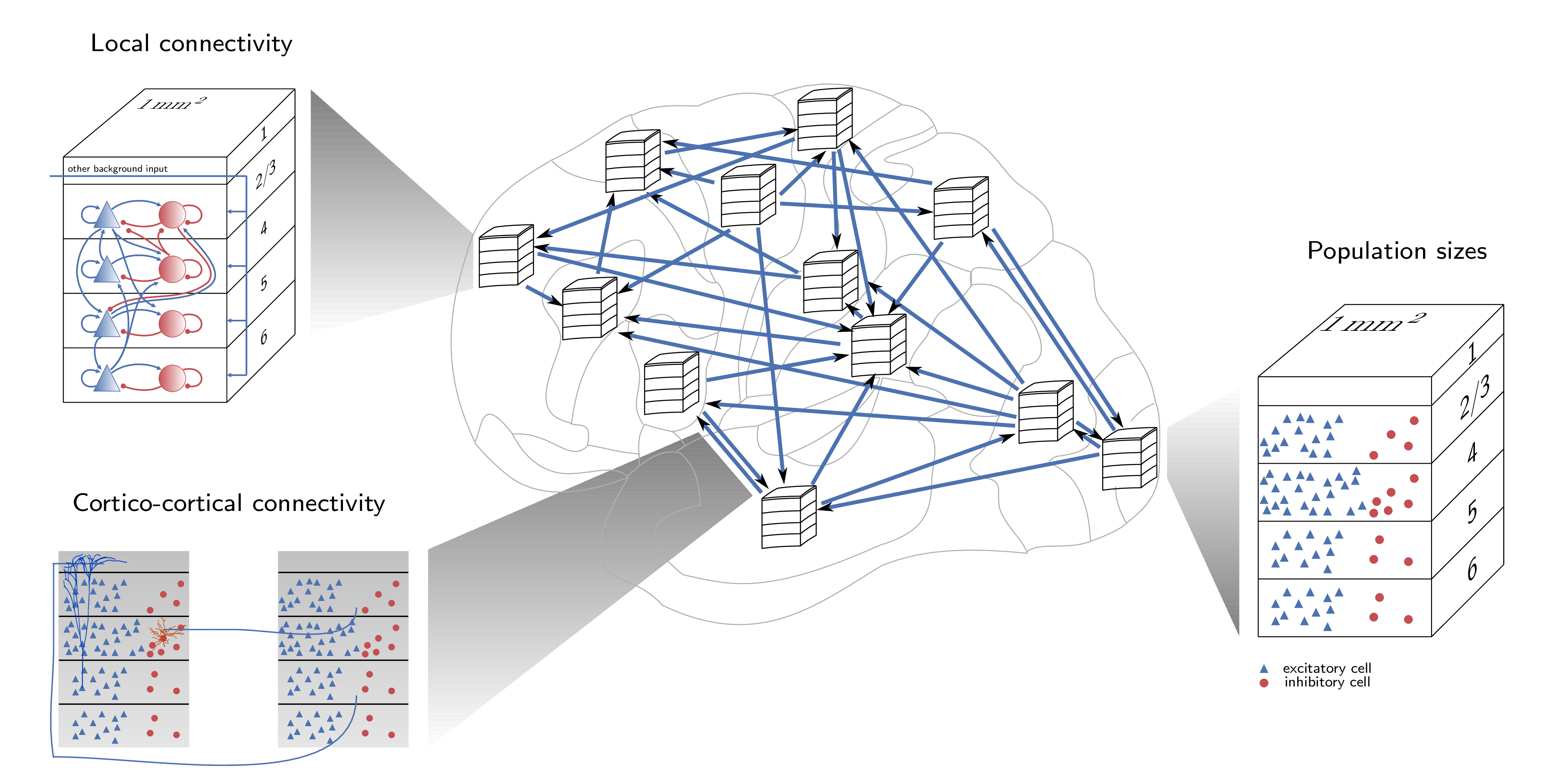
This repository holds the simulation data presented in Schmidt M, Bakker R, Shen K, Bezgin B, Diesmann M & van Albada SJ (2018) A multi-scale layer-resolved spiking network model of resting-state dynamics in macaque cortex. PLOS Computational Biology (accepted).

The simulation code of the model and the figure scripts for the paper reside in a public github repository available under the following url: https://github.com/INM-6/multi-area-model.
Simulations in this repository
This repository contains data from 17 different simulations that are presented in the paper. They are identified by their unique hashed labels that are generated from the parameter sets used in the respective simulation.
Here is a table that summarizes the simulations and their most important parameters:
| Connectivity matrix |
Coupling constant for cortico-cortical synaptic weights |
Increased external input onto population 5E and 6E |
Simulation time (ms) |
Figures in Schmidt et al. 2018 |
Simulation label |
| before stabilization |
1. |
1. |
10500 |
2 |
533d73357f |
| before stabilization |
1. |
1.125 |
10500 |
2 |
0adda4a542 |
| after stabilization |
1. |
1.125 |
10500 |
2,3,4,6,8 |
33fb595555 |
| after stabilization |
1.9 |
1.125 |
10500 |
5,6 |
3afaec94d6 |
| after stabilization |
1.9 |
1.125 |
100500 |
4,5,6,7,8,9 |
99c0024eac |
| after stabilization |
1.4 |
1.125 |
10500 |
8 |
783cedb0ff |
| after stabilization |
1.5 |
1.125 |
10500 |
8 |
380856f3b3 |
| after stabilization |
1.6 |
1.125 |
10500 |
8 |
5a7c6c2d6d |
| after stabilization |
1.7 |
1.125 |
10500 |
8 |
c1876856b1 |
| after stabilization |
1.75 |
1.125 |
10500 |
8 |
a30f6fba65 |
| after stabilization |
1.8 |
1.125 |
10500 |
4,8 |
1474e18844 |
| after stabilization |
2. |
1.125 |
10500 |
4,8 |
f18158895a |
| after stabilization |
2.1 |
1.125 |
10500 |
4,8 |
08a3a1a88c |
| after stabilization |
2.5 |
1.125 |
10500 |
4,6,8 |
5bdd72887b |
To reduce the required space, we here refrain from publishing the original spiking data (~150 GB) and publish only the post-processed data presented in the figures of Schmidt et al.
Citation
If you use these data, we ask you to cite our paper in your publication.
If you have questions regarding the code or scientific content, please create an issue on the simulation code repository on github: https://github.com/INM-6/multi-area-model.
Acknowledgements
This work was supported by the Helmholtz Portfolio Supercomputing and
Modeling for the Human Brain (SMHB), the European Union 7th Framework
Program (Grant 269921, BrainScaleS and 604102, Human Brain Project,
Ramp up phase) and European Unions Horizon 2020 research and
innovation program (Grant 720270, Human Brain Project, SGA1), the
Jülich Aachen Research Alliance (JARA), the Helmholtz young
investigator group VH-NG-1028,and the German Research Council (DFG
Grants SFB936/A1,Z1 and TRR169/A2) and computing time granted by the
JARA-HPC Ver- gabegremium and provided on the JARA-HPC Partition part
of the supercomputer JUQUEEN (Jülich Supercomputing Centre 2015) at
Forschungszentrum Jülich (VSR Computation Time Grant JINB33), and Priority
Program 2041 (SPP 2041) "Computational Connectomics" of the German Research
Foundation (DFG).

