|
|
@@ -1,3 +1,174 @@
|
|
|
-# geo-htseq-paper
|
|
|
+Working with model objects
|
|
|
+================
|
|
|
|
|
|
-A field-wide assessment of differential high throughput sequencing reveals widespread bias
|
|
|
+## Install
|
|
|
+
|
|
|
+- Download and install R <https://www.r-project.org> (and RStudio
|
|
|
+ <https://www.rstudio.com/products/rstudio/>).
|
|
|
+
|
|
|
+- Go to R console or open scripts/README.Rmd in RStudio.
|
|
|
+
|
|
|
+- Install these packages (this needs to be done once).
|
|
|
+
|
|
|
+``` r
|
|
|
+if(!require(brms)){
|
|
|
+ install.packages(c("brms", "here"))
|
|
|
+ library(brms)
|
|
|
+ library(here)
|
|
|
+}
|
|
|
+```
|
|
|
+
|
|
|
+## Run
|
|
|
+
|
|
|
+In R console OR in .Rmd file:
|
|
|
+
|
|
|
+- Load packages to R environment.
|
|
|
+
|
|
|
+``` r
|
|
|
+library(brms)
|
|
|
+library(here)
|
|
|
+```
|
|
|
+
|
|
|
+- Load the model object and print model summary.
|
|
|
+
|
|
|
+``` r
|
|
|
+m <- readRDS(here("models/anticons_detool.rds"))
|
|
|
+print(m)
|
|
|
+```
|
|
|
+
|
|
|
+ ## Family: bernoulli
|
|
|
+ ## Links: mu = logit
|
|
|
+ ## Formula: anticons ~ de_tool
|
|
|
+ ## Data: data (Number of observations: 2109)
|
|
|
+ ## Samples: 4 chains, each with iter = 2000; warmup = 1000; thin = 1;
|
|
|
+ ## total post-warmup samples = 4000
|
|
|
+ ##
|
|
|
+ ## Population-Level Effects:
|
|
|
+ ## Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
|
|
|
+ ## Intercept -3.51 0.23 -4.00 -3.09 1.00 942 1128
|
|
|
+ ## de_tooldeseq 2.66 0.25 2.19 3.18 1.00 1080 1153
|
|
|
+ ## de_tooledger 3.57 0.26 3.07 4.10 1.00 1051 1435
|
|
|
+ ## de_toollimma 3.58 0.42 2.77 4.42 1.00 1693 2093
|
|
|
+ ## de_toolunknown 2.46 0.25 1.99 2.98 1.00 1052 1153
|
|
|
+ ##
|
|
|
+ ## Samples were drawn using sampling(NUTS). For each parameter, Bulk_ESS
|
|
|
+ ## and Tail_ESS are effective sample size measures, and Rhat is the potential
|
|
|
+ ## scale reduction factor on split chains (at convergence, Rhat = 1).
|
|
|
+
|
|
|
+- Get fitted coefficients with 95% credible intervals.
|
|
|
+
|
|
|
+``` r
|
|
|
+posterior_summary(m)
|
|
|
+```
|
|
|
+
|
|
|
+ ## Estimate Est.Error Q2.5 Q97.5
|
|
|
+ ## b_Intercept -3.514582 0.2300317 -4.001753 -3.088126
|
|
|
+ ## b_de_tooldeseq 2.656732 0.2492621 2.188271 3.183202
|
|
|
+ ## b_de_tooledger 3.571890 0.2642955 3.074863 4.103799
|
|
|
+ ## b_de_toollimma 3.579595 0.4163380 2.773039 4.424586
|
|
|
+ ## b_de_toolunknown 2.461885 0.2476724 1.990004 2.975438
|
|
|
+ ## lp__ -969.443536 1.6153116 -973.461001 -967.323674
|
|
|
+
|
|
|
+- Get the full posterior.
|
|
|
+
|
|
|
+``` r
|
|
|
+post <- posterior_samples(m)
|
|
|
+head(post)
|
|
|
+```
|
|
|
+
|
|
|
+ ## b_Intercept b_de_tooldeseq b_de_tooledger b_de_toollimma b_de_toolunknown lp__
|
|
|
+ ## 1 -3.300792 2.355140 3.418613 3.433826 2.191998 -968.0241
|
|
|
+ ## 2 -3.485360 2.606697 3.559653 3.530865 2.329601 -967.5360
|
|
|
+ ## 3 -3.628822 2.705505 3.763745 3.932852 2.560540 -967.7857
|
|
|
+ ## 4 -3.621123 2.917484 3.643992 3.647139 2.545874 -968.5439
|
|
|
+ ## 5 -3.795040 2.929855 3.797475 3.520305 2.633857 -968.9733
|
|
|
+ ## 6 -3.248306 2.532679 3.443725 3.528865 2.130235 -969.7214
|
|
|
+
|
|
|
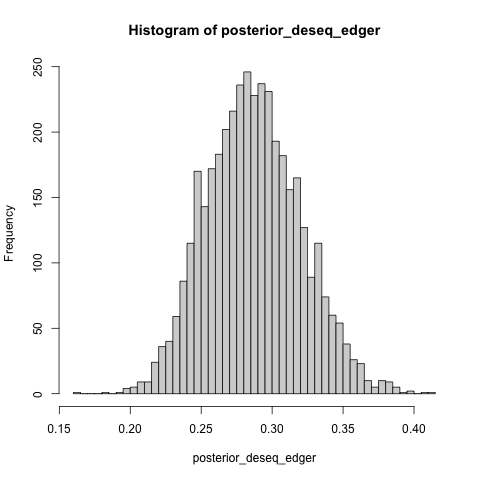
+- What is the estimated difference in the proportion of
|
|
|
+ anti-conservative p value histograms between DESeq2 and EdgeR?
|
|
|
+
|
|
|
+``` r
|
|
|
+posterior_deseq_edger <- inv_logit_scaled(post$b_de_tooldeseq - post$b_de_tooledger)
|
|
|
+hist(posterior_deseq_edger, breaks = 40)
|
|
|
+```
|
|
|
+
|
|
|
+
|
|
|
+
|
|
|
+- The posterior summary for the effect size.
|
|
|
+
|
|
|
+``` r
|
|
|
+posterior_summary(posterior_deseq_edger)
|
|
|
+```
|
|
|
+
|
|
|
+ ## Estimate Est.Error Q2.5 Q97.5
|
|
|
+ ## [1,] 0.2870903 0.03315653 0.2267546 0.3541821
|
|
|
+
|
|
|
+The estimated effect size is somewhere between 23% and 35%.
|
|
|
+
|
|
|
+- Get data from model object.
|
|
|
+
|
|
|
+``` r
|
|
|
+data <- m$data
|
|
|
+head(data)
|
|
|
+```
|
|
|
+
|
|
|
+ ## anticons de_tool
|
|
|
+ ## 1 1 unknown
|
|
|
+ ## 2 0 unknown
|
|
|
+ ## 3 1 edger
|
|
|
+ ## 4 0 cuffdiff
|
|
|
+ ## 5 0 limma
|
|
|
+ ## 6 0 unknown
|
|
|
+
|
|
|
+- Extract stan code from model object. This is the fullest model
|
|
|
+ description.
|
|
|
+
|
|
|
+``` r
|
|
|
+stancode(m)
|
|
|
+```
|
|
|
+
|
|
|
+ ## // generated with brms 2.15.0
|
|
|
+ ## functions {
|
|
|
+ ## }
|
|
|
+ ## data {
|
|
|
+ ## int<lower=1> N; // total number of observations
|
|
|
+ ## int Y[N]; // response variable
|
|
|
+ ## int<lower=1> K; // number of population-level effects
|
|
|
+ ## matrix[N, K] X; // population-level design matrix
|
|
|
+ ## int prior_only; // should the likelihood be ignored?
|
|
|
+ ## }
|
|
|
+ ## transformed data {
|
|
|
+ ## int Kc = K - 1;
|
|
|
+ ## matrix[N, Kc] Xc; // centered version of X without an intercept
|
|
|
+ ## vector[Kc] means_X; // column means of X before centering
|
|
|
+ ## for (i in 2:K) {
|
|
|
+ ## means_X[i - 1] = mean(X[, i]);
|
|
|
+ ## Xc[, i - 1] = X[, i] - means_X[i - 1];
|
|
|
+ ## }
|
|
|
+ ## }
|
|
|
+ ## parameters {
|
|
|
+ ## vector[Kc] b; // population-level effects
|
|
|
+ ## real Intercept; // temporary intercept for centered predictors
|
|
|
+ ## }
|
|
|
+ ## transformed parameters {
|
|
|
+ ## }
|
|
|
+ ## model {
|
|
|
+ ## // likelihood including constants
|
|
|
+ ## if (!prior_only) {
|
|
|
+ ## target += bernoulli_logit_glm_lpmf(Y | Xc, Intercept, b);
|
|
|
+ ## }
|
|
|
+ ## // priors including constants
|
|
|
+ ## target += student_t_lpdf(Intercept | 3, 0, 2.5);
|
|
|
+ ## }
|
|
|
+ ## generated quantities {
|
|
|
+ ## // actual population-level intercept
|
|
|
+ ## real b_Intercept = Intercept - dot_product(means_X, b);
|
|
|
+ ## }
|
|
|
+
|
|
|
+## This document
|
|
|
+
|
|
|
+This README.md was generated by running:
|
|
|
+
|
|
|
+``` r
|
|
|
+rmarkdown::render("scripts/README.Rmd", output_file = here::here("README.md"))
|
|
|
+```
|